USCLobes Atlas
A high-resolution anatomical MRI atlas with parcellation into lobes
Developed by Anand A. Joshi, Soyoung Choi, Jessica L. Wisnowski, Justin P. Haldar, David W. Shattuck, Hanna Damasio, and Richard M. Leahy

This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License
When using the USCLobes Atlas in your published work, please cite the following reference paper:
Joshi AA, Choi S, Chong M, Sonkar G, Gonzalez-Martinez J, Nair D, Wisnowski JL, Haldar JP, Shattuck DW, Damasio H, Leahy RM (2022) A Hybrid High-Resolution Anatomical MRI Atlas with Sub-parcellation of Cortical Gyri using Resting fMRI. Journal of Neuroscience Methods 374:109566.
Atlas Description
The USCLobes atlas segments the brain into larger regions than those provided by the default BrainSuite atlases. The detailed description of the atlas development can be found in the description page of the BCI-DNI_brain atlas description.
Anatomical Parcellation
A high-resolution 3D MPRAGE scan (TE=4.33 ms; TR=2070 ms; TI=1100ms; flip angle=12 degrees; resolution=0.547×0.547×0.802 mm) was acquired on a 3T Siemens MAGNETOM Tim Trio using a 32-channel head coil. Data were acquired 5 times and averaged. The subject is a typical right-handed woman in her mid-thirties with a brachiocephalic brain and no rare anomalies.
Preprocessing and labeling of the MPRAGE image was performed using BrainSuite [1,2]. This MRI volume was used to generate the BCI-DNI atlas as described here.
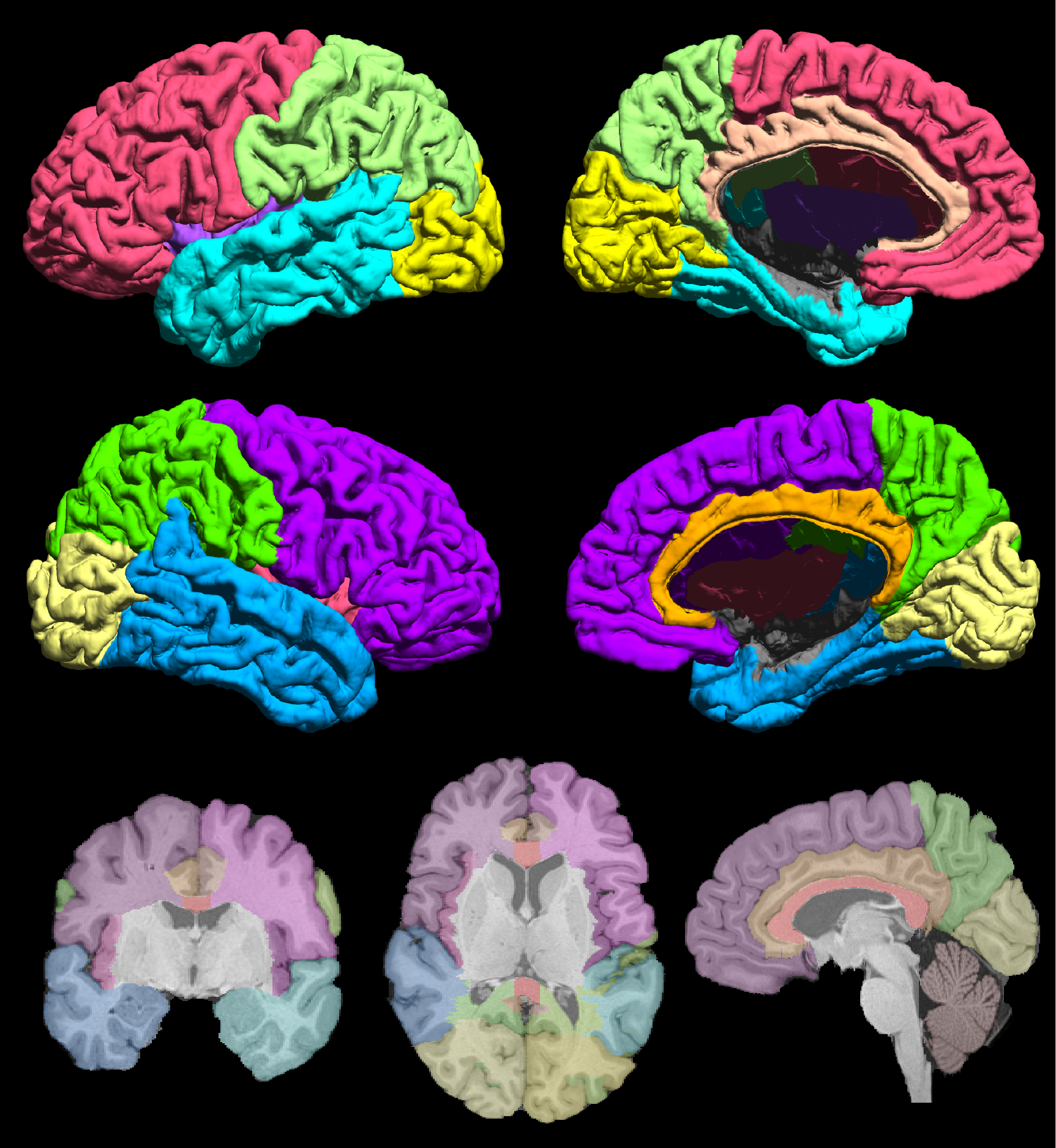
Fig. 1 The USCLobes atlas with 15 regions of interest.
Regions of Interest
15 ROIs are delineated on the volumetric labels of the atlas. 12 of these regions are labeled on the surface.
-
(L/R) Frontal Lobe
(L/R) Parietal Lobe
(L/R) Temporal Lobe
(L/R) Occipital Lobe
(L/R) Insula
(L/R) Cingulate
Brainstem
Cerebellum
Corpus Callosum
Usage
Detailed instructions for using the USCLobes atlas with BrainSuite, FreeSurfer, and FSL are available here.
References
1. Damasio, H., (1995), ‘Human brain anatomy in computerized images’, Oxford university press.
2. Joshi, A., (2012), ‘A method for automated cortical surface registration and labeling’, International Workshop on Biomedical Image Registration, pp 180-189.