4. The BrainSuite Pipelines
BrainSuite provides pipelines for analyzing Anatomical T1-weighted MRI, Diffusion MRI, and Functional MRI. The BrainSuite BIDS App provides the capability of running each of these on datasets stored in the BIDS format. Below are the details of the pipelines.
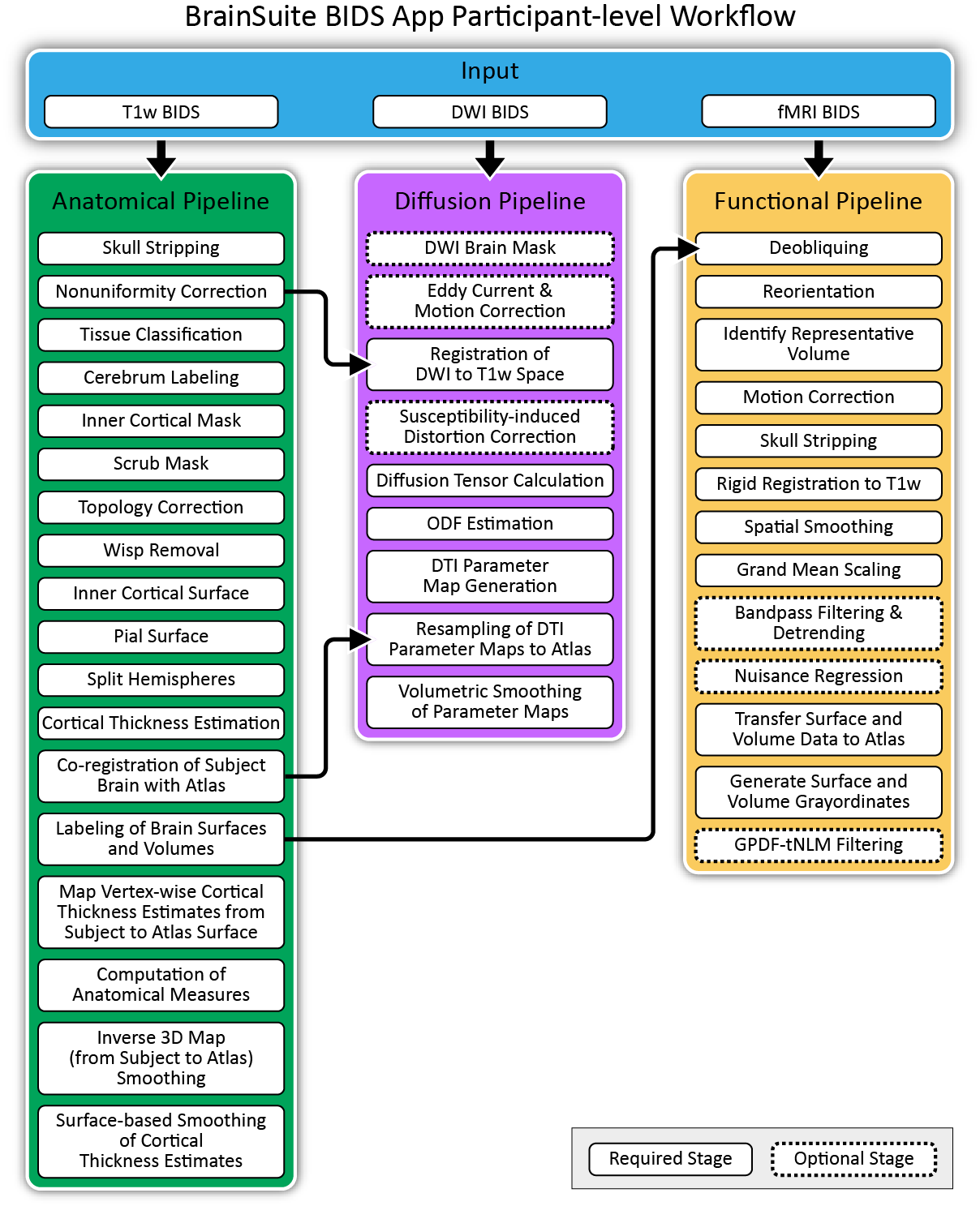
Fig 4. BrainSuite Participant-level Workflows.
4.1. Anatomical Pipeline
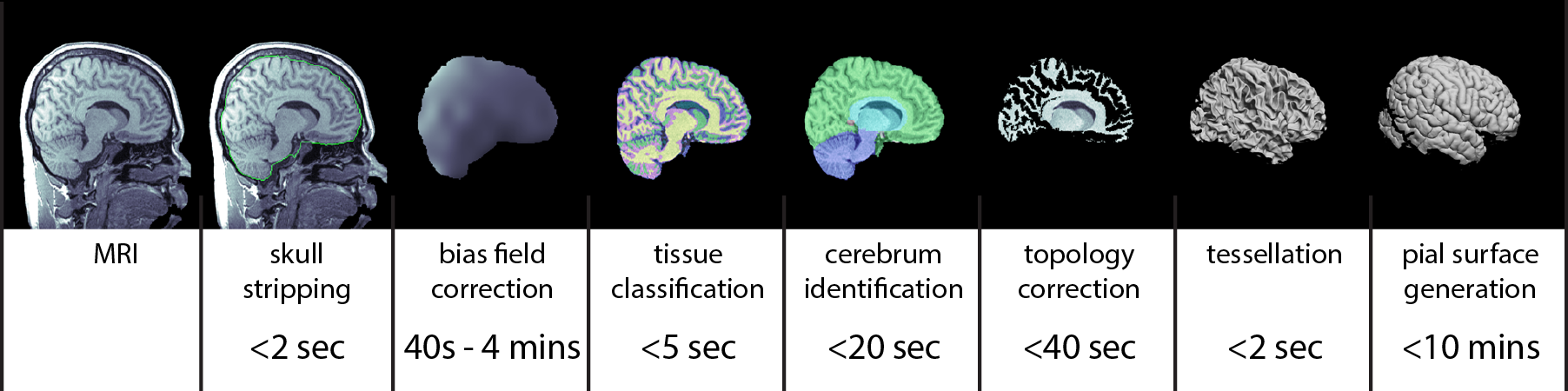
The BrainSuite Anatomical Pipeline (BAP) processes T1-weighted (T1w) MRI to generate brain surfaces and volumes that are consistently registered and labeled according to a reference anatomical atlas. The major stages in BAP comprise:
Cortical surface extraction (CSE), which includes skull-stripping, tissue classification, gross labeling of brain structures, and modeling of the inner and outer boundaries of the cerebral cortex.
Cortical thickness estimation based on partial volume estimates and the anisotropic diffusion equation.
Surface-constrained volumetric registration (SVReg) to generate a mapping to a labeled reference atlas and label the cortical surface and brain volume.
Mapping of cortical thickness estimates to the atlas space for group-level analysis.
Computation of subject-level statistics (e.g., mean GM volume within ROIs, cortical thickness within surface ROIs).
Smoothing of Jacobian determinants maps (magnitudes of the 3D deformation fields of MRI images registered to the atlas) in preparation for tensor-based morphometry analysis. *
* This step is not part of the BrainSuite software’s BAP workflow and only exists in the BrainSuite BIDS App’s BAP workflow.
4.2. Diffusion Pipeline
The Diffusion Pipeline is based on BDP and performs several steps to process diffusion MRI. These include:
Eddy current and subject motion correction using FSL’s eddy. **
Processing with BDP to perform: * Processing of diffusion weighted imaging (DWI) to correct geometric image distortion (based on either field maps or nonlinear registration to a corresponding T1-weighted MRI). * Coregistration of the DWI to the T1w scan. * Fitting of diffusion tensor models to the DWI data. * Fitting of orientation distribution functions to the DWI data (using FRT, FRACT, GQI, 3D-SHORE, or ERFO as appropriate). * Computation of diffusion indices (FA, MD, AxD, RD, GFA).
Mapping of diffusion parametric maps to atlas space for group-level analysis. **
Smoothing of diffusion parametric maps in preparation for voxel-wise group-level analyses. **
** These steps are not part of the standalone command-line BDP program and have been added for the BrainSuite BIDS App’s Diffusion Pipeline.
4.3. Functional Pipeline
The BrainSuite Functional Pipeline (BFP) processes resting-state and task-based fMRI data. BFP processes 4D fMRI datasets using a combination of tools from AFNI, FSL, BrainSuite and additional in-house tools developed for BrainSuite:
Performs deobliquing and reorientation using tools from AFNI.
Identifies a representative fMRI volume for use in motion correction and registration to the T1w data.
Corrects for motion by applying a 5-parameter rigid registration of each time point volume in the fMRI data to the reference volume using tools from AFNI.
Detects outliers by either computing structural similarity index (SSIM) between each fMRI time point and the reference image or by using DVARS from FSL.
Registers the fMRI data to the corresponding T1w anatomical data.
Skull strips fMRI data using T1w brain mask.
Applies spatial smoothing, grand-mean scaling, temporal bandpass filtering, and detrending.
Performs nuisance signal regression, which regresses signals from cerebrospinal fluid, white matter, and motion using the FEAT model in FSL.
Generates a representation of the fMRI data in atlas space and grayordinate space in preparation for group-level analysis.
Applies a global PDF-based nonlocal means filter, a data-driven approach for filtering fMRI signals based on the probability density functions of the correlations between time series across spatial points throughout the brain [Li2020].