Inner Cortical Surface
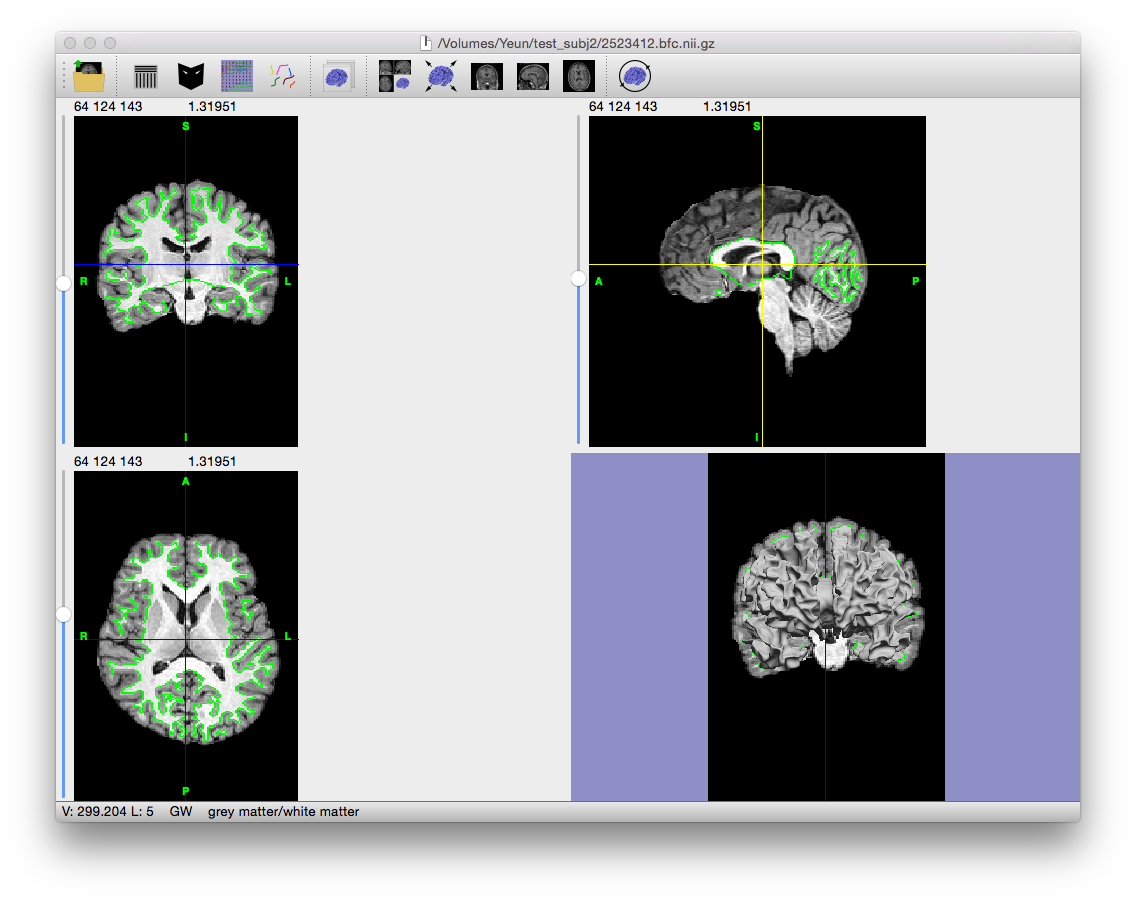
 This module generates a surface mesh based on the cleaned and corrected inner cortical boundary mask.
This module generates a surface mesh based on the cleaned and corrected inner cortical boundary mask.
If adjusting the parameters of this stage does not produce a satisfactory surface, rerun the stages that created the mask (cortex mask selection, scrub mask, topology correction, and wisp removal) and change the parameters to achieve a better foundation mask with which to generate the surface.
Parameters
- Smoothing iterations (typical range is 5–20)
- Number of smoothing iterations to apply.
- Smoothing constant (typical range is 0.1–1)
- The strength of the smoothing, ranging from 0 (no smoothing) to 1 (most smoothing).
- Curvature weight (typical range is 1–5)
- Adjusts the smoothing based on a measure of local curvature
Command-Line Usage
dfs: generates mesh surfaces using an isosurface algorithm.
usage: dfs -i input -o output [optional settings]
example: dfs -i input_mri.nii.gz -o surface.dfs
Required Settings:
| Flags | Description |
|---|---|
-i <input volume> |
input 3D volume |
-o <output surface> |
output surface mesh file |
Optional Settings:
| Flags | Description |
|---|---|
-c <image volume> |
shade surface model with data from image volume; a set of vertex colors is generated within the file |
-n <value> |
number of smoothing iterations [default: 10] |
-a <value> |
smoothing constant [default: 0.5] |
-w <value> |
curvature weighting [default: 0] |
-f <value> |
scaling percentile [default: 0.999] |
-nz |
tessellate non-zero voxels |
-gt <threshold> |
tessellate voxels greater than <threshold> |
-lt <threshold> |
tessellate voxels less than <threshold> |
-eq <value> |
tessellate voxels equal to <value> |
-z |
zero-pad volume (avoids clipping at edges) |
--nonormals |
do not compute vertex normals |
--postsmooth |
smooth vertices after coloring |
-v <level> |
verbosity (0 = quiet) [default: 1] |
--timer |
timing function |
Example Result
Output
If “save output of each stage automatically” is checked on the Cortical Surface Extraction sequence, the following files are generated (where filename_prefix is the filename of the MRI scan without the file extension, e.g. “testsubj” for the file “testsubj.nii”):
| Filename | Contents |
|---|---|
| filename_prefix.inner.cortex.dfs | Surface representing the boundary between white matter and cortical grey matter. Provides the foundation for the generation of the pial surface in the next step. |
Restore from Previous Session
If BrainSuite was interrupted while performing this stage or to change the parameters for this stage and rerun after fully processing a scan, click “Restore From Previous…” on the bottom of this stage’s dialog box and load the original MRI scan. BrainSuite will automatically load all of the files generated in previous stages, allowing processing to restart from this intermediate stage.